Deep Embedding for Single-cell Clustering (DESC)
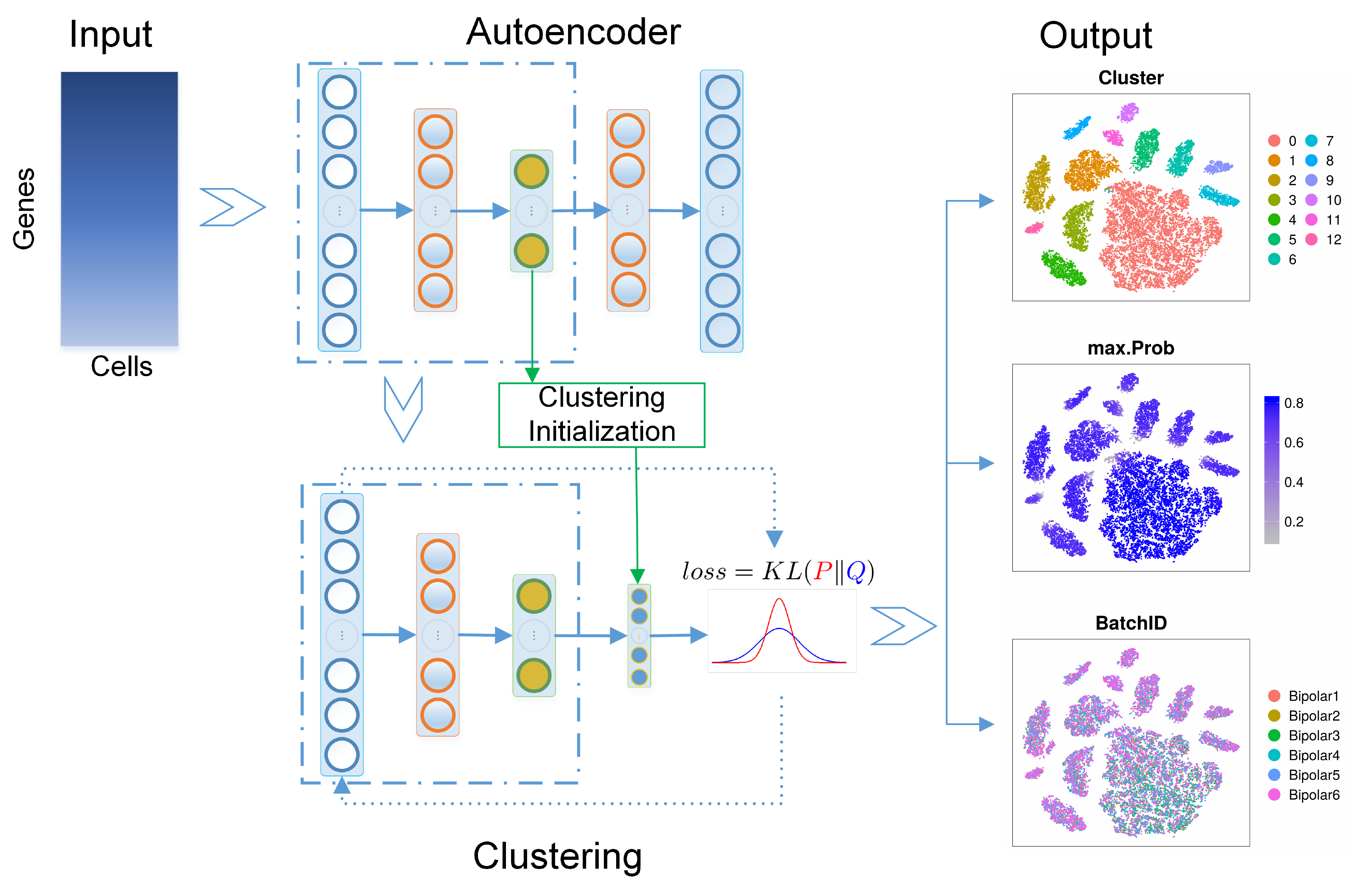
DESC is an unsupervised deep learning algorithm for clustering scRNA-seq data. The algorithm constructs a non-linear mapping function from the original scRNA-seq data space to a low-dimensional feature space by iteratively learning cluster-specific gene expression representation and cluster assignment based on a deep neural network. This iterative procedure moves each cell to its nearest cluster, balances biological and technical differences between clusters, and reduces the influence of batch effect. DESC also enables soft clustering by assigning cluster-specific probabilities to each cell, which facilitates the identification of cells clustered with high-confidence and interpretation of results.
For thorough details, see our paper: https://www.nature.com/articles/s41467-020-15851-3
Usage
The desc package is an implementation of deep embedding for single-cell clustering. With desc, you can:
- Preprocess single cell gene expression data from various formats.
- Build a low-dimensional representation of the single-cell gene expression data.
- Obtain soft-clustering assignments of cells.
- Visualize the cell clustering results and the gene expression patterns.
Because of the difference between tensorflow 1* and tensorflow 2*, we updated our desc algorithm into two version such that it can be compatible with tensorflow 1* and tensorflow 2*, respectively.
- For
tensorflow 1*, we releaseddesc(2.0.3). Please see our jupyter notebook example desc_2.0.3_paul.ipynb - For
tensorflow 2*, we releaseddesc(2.1.1). Please see our jupyter notebook example desc_2.1.1_paul.ipynb
References
Please consider citing the following reference:
- Xiangjie Li, Kui Wang, Yafei Lyu, Huize Pan, Jingxiao Zhang, Dwight Stambolian, Katalin Susztak, Muredach P. Reilly, Gang Hu, Mingyao Li. Deep learning enables accurate clustering and batch effect removal in single-cell RNA-seq analysis. Nature Communication 11, 2338 (2020). https://www.nature.com/articles/s41467-020-15851-3